All examples shown here were recorded with light-sheet or confocal microscopy and then deconvolved with NovoDeblur using a computed point spread function (PSF).
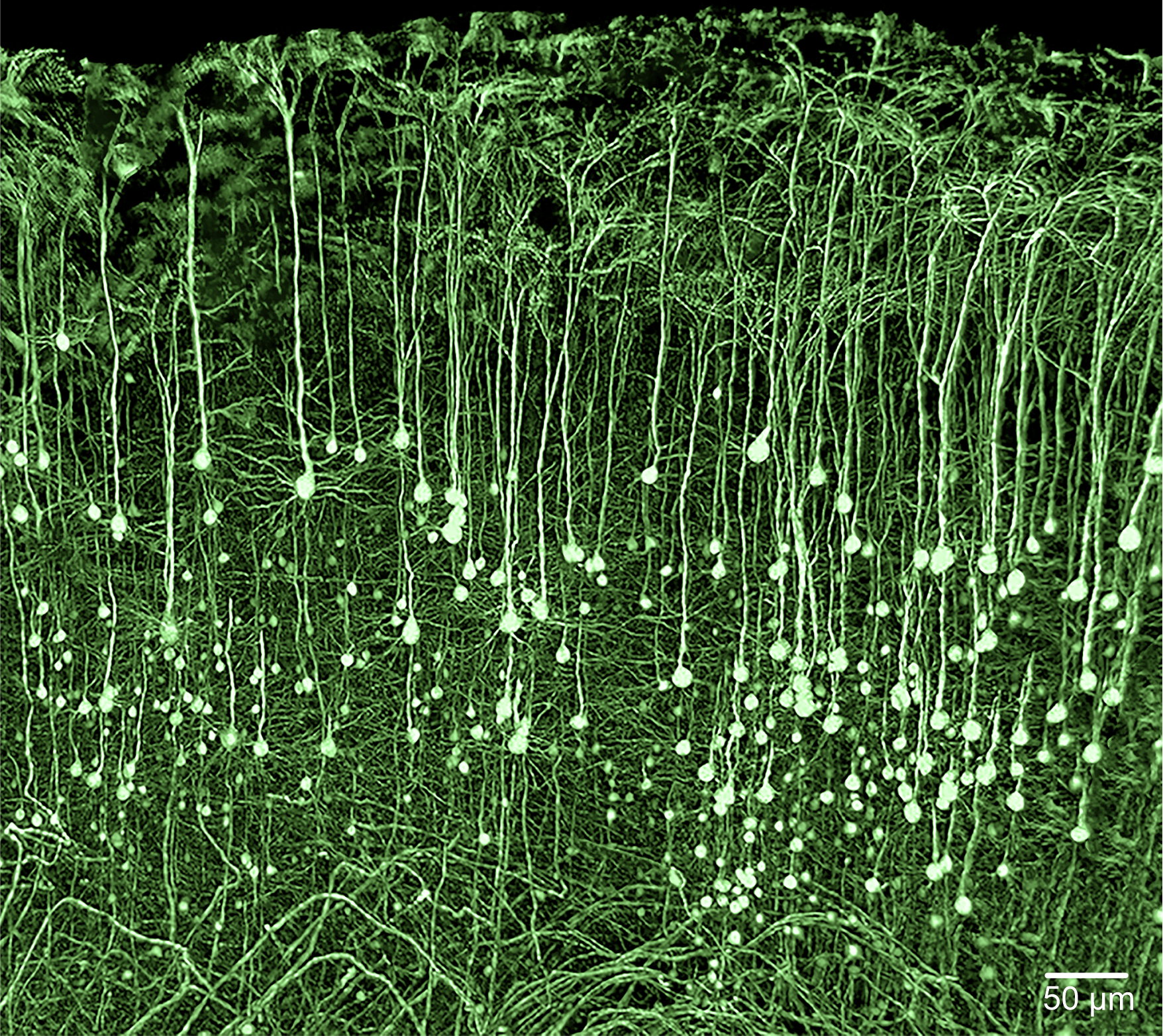
3D reconstruction of GFP-expressing pyramidal neurons in the hippocampus of a three-week-old mouse, acquired using light-sheet microscopy. Imaging was performed with a 4x objective (Olympus XL FLUOR 4x, NA 0.28) and a 0.63-fold post-demagnification. Excitation wavelength was 488 nm, and emission was detected at 520 nm. The imaging medium had a refractive index of 1.45. Raw data were deconvolved using NovoDeblur prior to 3D reconstruction.
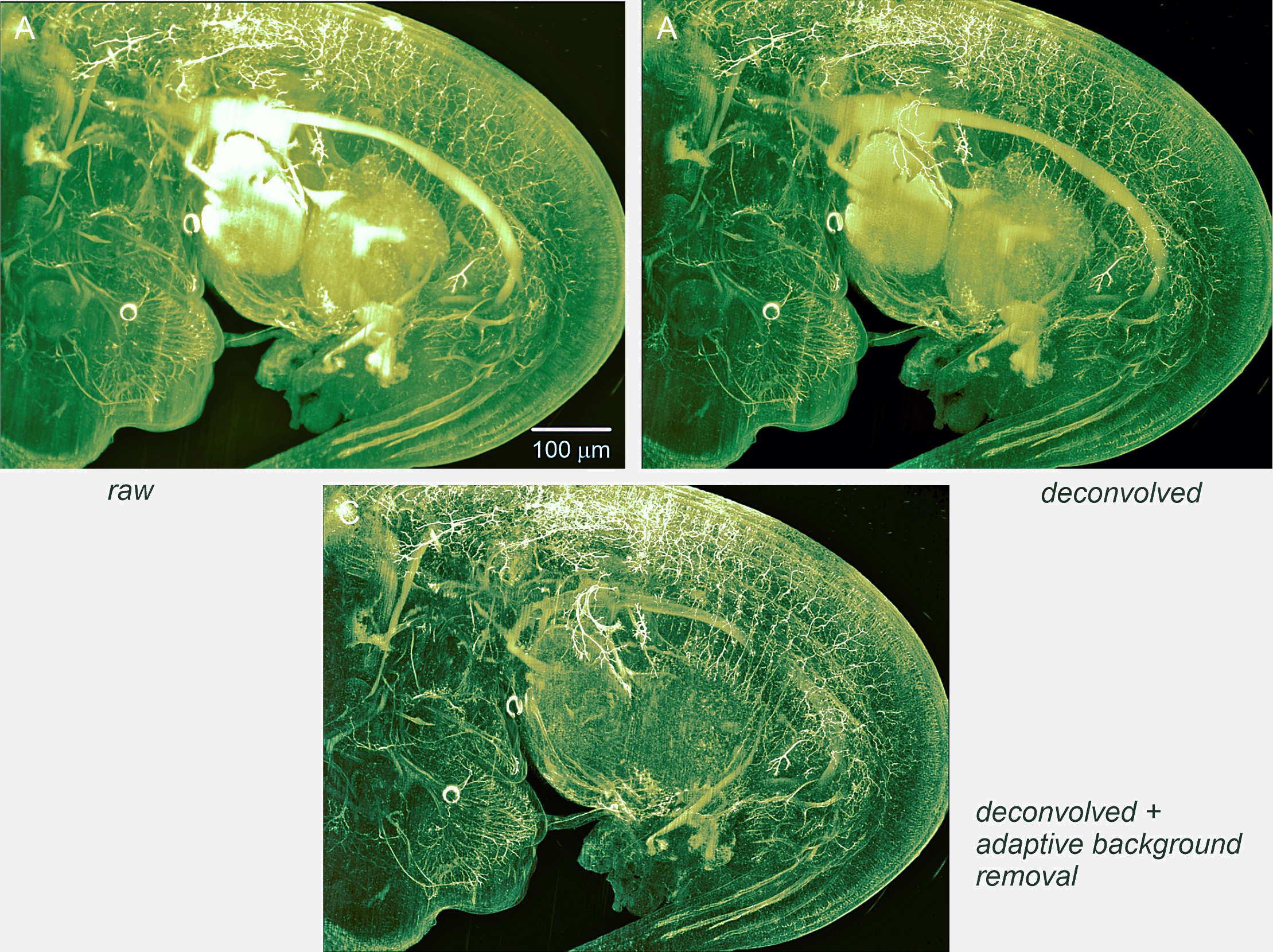
Visualization of nerve fibers highlighted by NF-160 fluorescence labeling using light-sheet microscopy. Imaging was performed with a cylindrical lens of 80 mm focal length and a slit aperture 6 mm wide (objective: Zeiss FLUAR 2.5x, NA 0.12, 0.5x post-magnification). A) Raw data: 3D reconstructions were generated from 668 slices, each with a resolution of 1392 x 1040 pixels, captured with a Cool Snap Cf CCD camera (Roper Scientific, Germany). The illumination time for each slice was 430 ms. B) Deconvolution without background subtraction: Deconvolution was carried out with the following parameters: NA = 0.12, excitation wavelength = 488 nm, emission wavelength = 520 nm, refractive index = 1.561, light-sheet NA = 0.05, stop criterion = 0.5%, maximum iterations = 100, histogram clipping = 0.01, and no damping. Total processing time was under 5 minutes on a 16-core workstation with 256 GB RAM and an NVIDIA P6000 GPU. C) Deconvolution with adaptive background subtraction: The same parameters as in B) were used, with rolling-ball background subtraction (75 µm filter radius) applied before deconvolution.
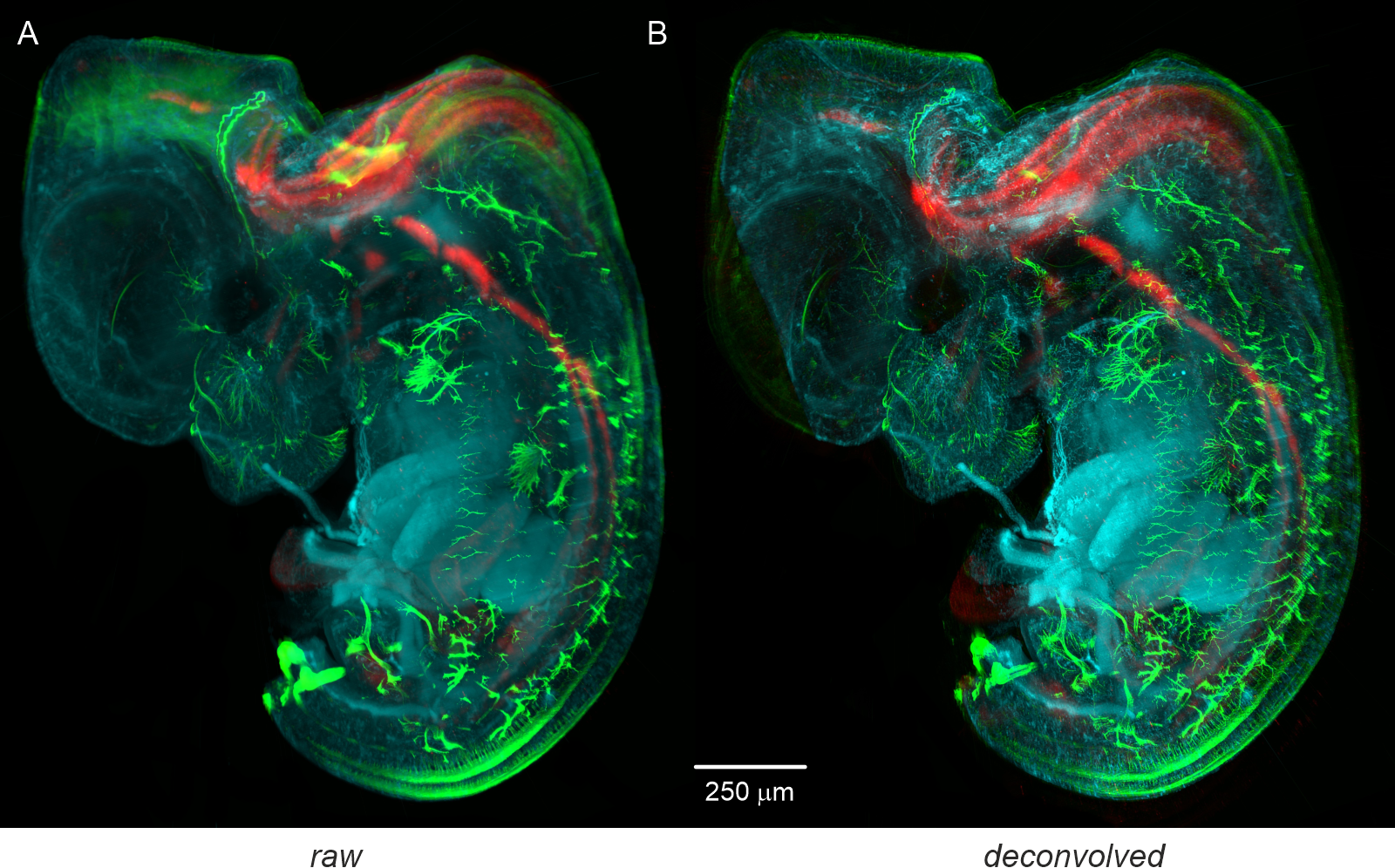
The fluorophores used for immunostaining were GFP (excitation wavelength = 488 nm), 2H3 (excitation wavelength = 550 nm), and Phox2B (excitation wavelength = 647 nm). Imaging was performed with a light-sheet microscope equipped with a high-Rayleigh-range light-sheet generator (NA = 0.05) and an objective lens (Olympus XL Fluor 4x, NA 0.28, 0.63x post-magnification). A) Original data: Maximum intensity projection (MIP) generated from 600 slices, each with a resolution of 2560 x 2160 pixels, recorded with an AndorNeo CCD camera (Oxford Instruments, Germany). B) Deconvolved data: MIP generated from the deconvolved data. Deconvolution was performed with the following parameters: NA = 0.28, excitation wavelengths = {488 nm, 550 nm, 647 nm}, emission wavelengths = {520 nm, 570 nm, 695 nm}, refractive index = 1.561, light-sheet NA = 0.05, stop criterion = 0.5%, maximum iterations = 100, histogram clipping = 0.01, no regularization, and a rolling-ball filter radius of 40 µm.

Imaging was performed with a light-sheet microscope equipped with a high-Rayleigh-range light-sheet generator (NA = 0.05) and a 4x objective lens (Olympus XL FLUOR 4x, NA 0.28, 2x post-magnification). A) Original data: Maximum intensity projection (MIP) generated from 777 slices, each with a resolution of 2560 x 2160 pixels, recorded with an AndorNeo CCD camera (Oxford Instruments, Germany). B) Deconvolved data: Deconvolution was performed with the following parameters: NA = 0.28, excitation wavelength = 488 nm, emission wavelength = 520 nm, refractive index = 1.561, light-sheet NA = 0.05, stop criterion = 0.5%, maximum iterations = 100, histogram clipping = 0.01, with no regularization and no background subtraction.
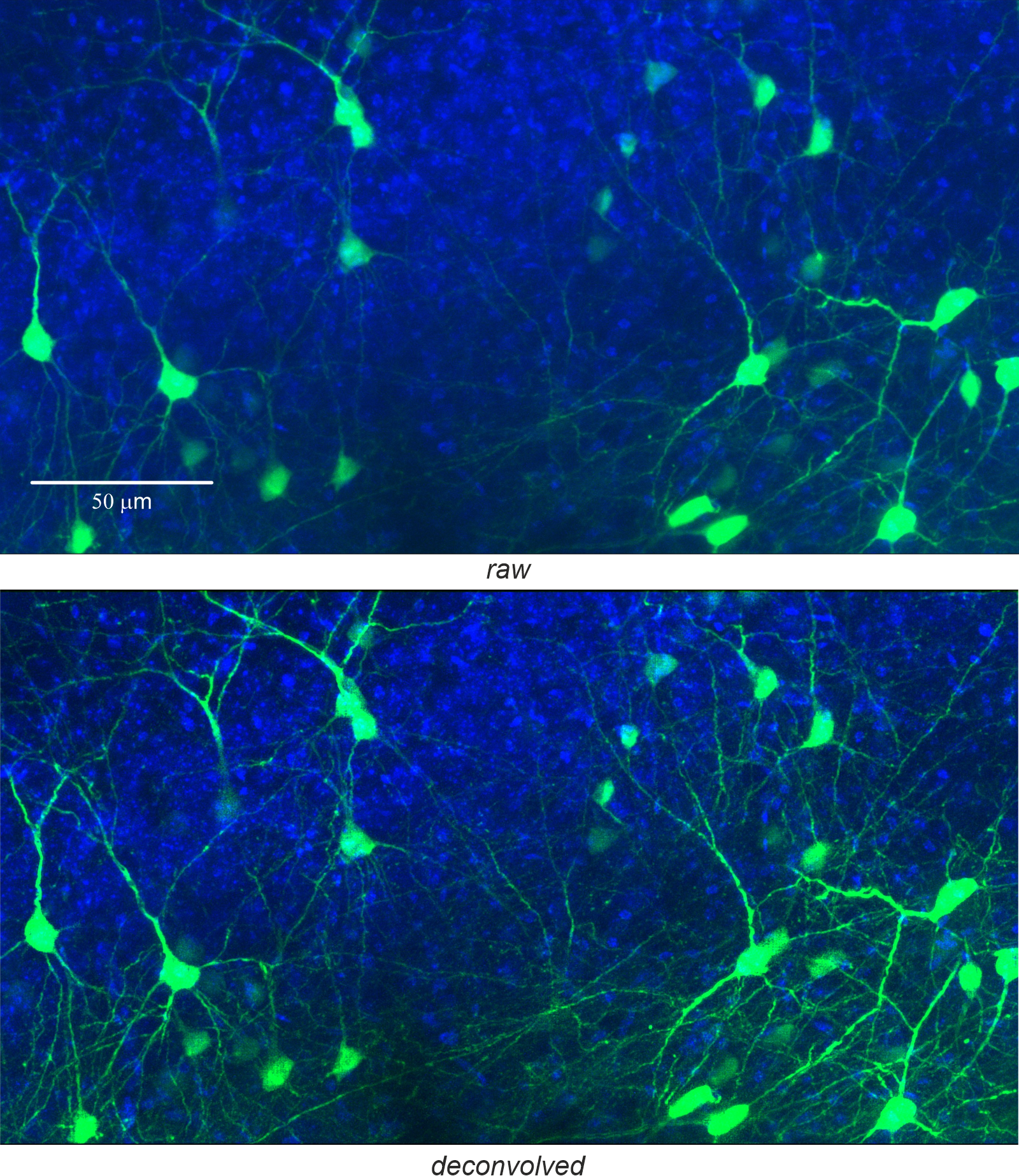
Objective: Zeiss 60x, NA 0.8; excitation wavelength = 488 nm; emission wavelength = 520 nm; refractive index = 1.45. A) Original data: MIP generated from 117 slices. B) Deconvolved data: Deconvolution parameters were NA = 0.8, excitation wavelength = 488 nm, emission wavelength = 520 nm, refractive index = 1.45, pinhole diameter = 30 µm, stop criterion = 0.1%, maximum iterations = 100, histogram clipping = 0.01, regularization = 40%, and rolling-ball filter radius = 2 µm.
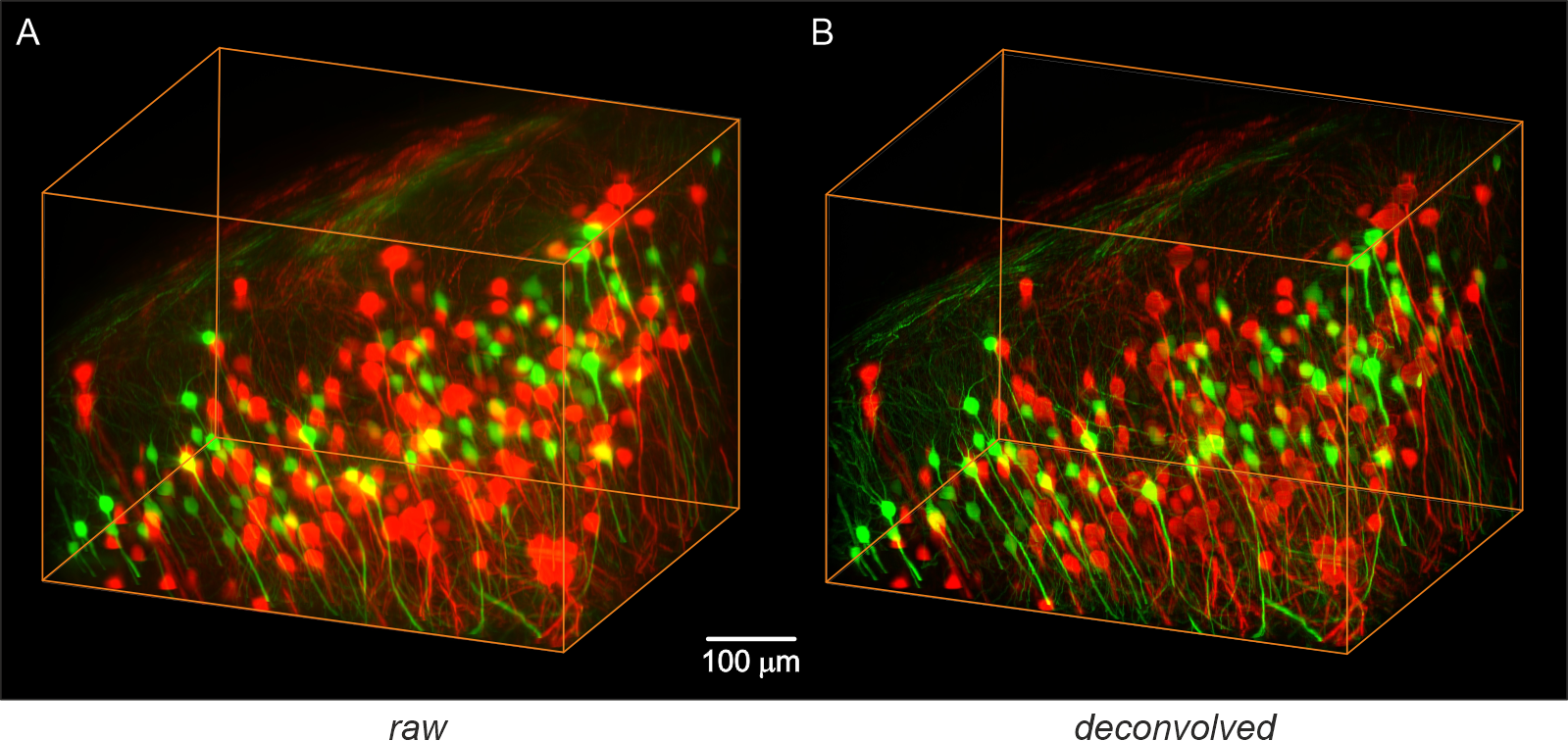
Imaging was performed with a light-sheet microscope equipped with a high-Rayleigh-range light-sheet generator (NA = 0.05) and an Olympus 25x objective (NA = 1.0). A) Original data: Maximum intensity projection (MIP) generated from 2 x 786 slices, each with a resolution of 2560 x 2160 pixels, recorded with an AndorNeo CCD camera (Oxford Instruments, Germany). B) Deconvolved data: The same dataset after deconvolution. Deconvolution parameters were NA = 1.0, excitation wavelengths = {488 nm, 532 nm}, emission wavelengths = {520 nm, 560 nm}, refractive index = 1.561, light-sheet NA = 0.05, stop criterion = 0.5%, maximum iterations = 100, histogram clipping = 0.01, no damping, and a rolling-ball filter radius of 15 µm.
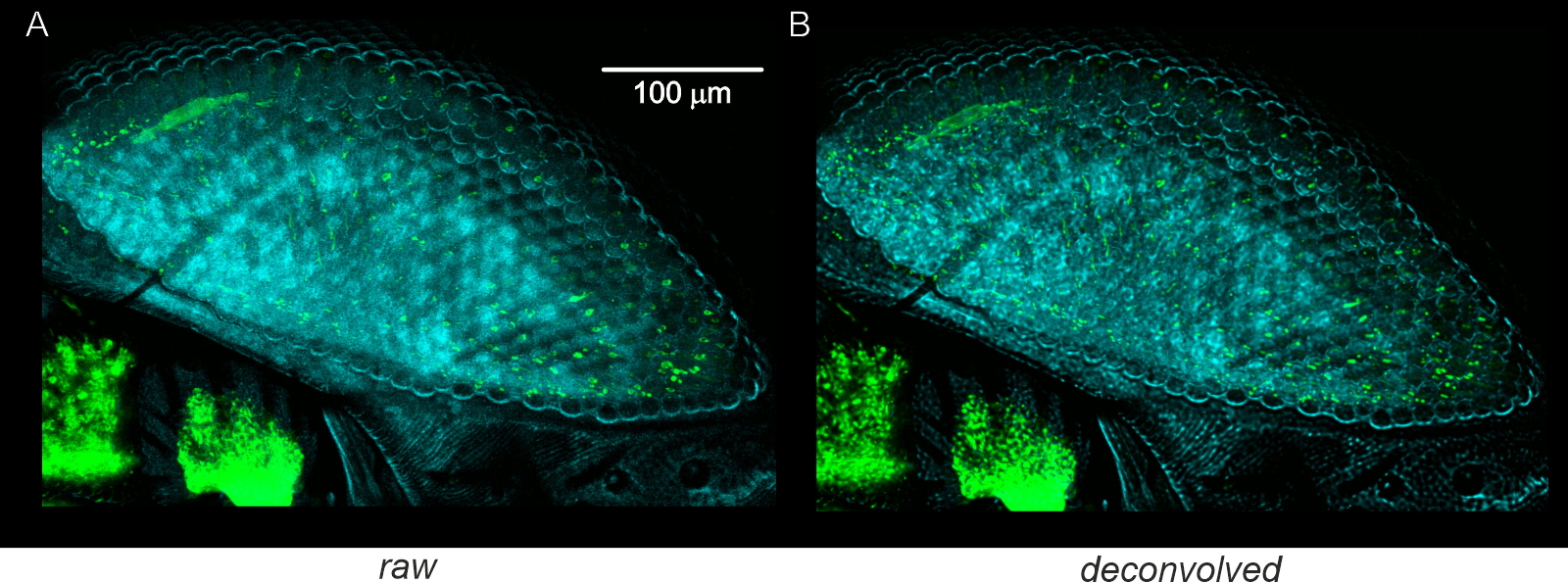
A) Original data: Confocal microscopy reconstructions of the faceted eye of a fruit fly using neuronal staining and autofluorescence imaging (objective: 25x, NA = 0.7). Maximum intensity projection (MIP) generated from 2 x 69 slices. B) Deconvolved data: MIP generated from the deconvolved data (69 slices with a resolution of 1024 x 1024 pixels). Deconvolution parameters were NA = 0.8, excitation wavelength = 488 nm, emission wavelength = 520 nm, refractive index = 1.45, pinhole diameter = 30 µm, stop criterion = 0.1%, maximum iterations = 100, histogram clipping = 0.01, 10% regularization, and no background subtraction.